Phenotype MicroArrays for Mammalian Cells
Phenotype MicroArray Mammalian assays are cell-based assays used to investigate up to 1,400 metabolic and chemical sensitivity phenotypes of mammalian cells. They use 96 well plates pre-loaded with carbon-energy and nitrogen substrates, ions, hormones/cytokines and anti-cancer agents. Plates are inoculated with the cells of interest and Biolog’s proprietary redox dye. In some wells, the cells are stimulated and in other wells inhibited. The generation of energy-rich NADH by the cells reduces the redox dye and brings about a color change which is then read with Biolog’s OmniLog automated incubator-reader.
Phenotype MicroArray Mammalian Assays can be used for diverse research and applications including:
- Correlating genotypes with phenotypes
- Studying metabolic reprogramming in cancer and anti-cancer drug sensitivity
- Drug discovery research and toxicity screening
- Cell line and bioprocess improvement
- Cellular metabolism, metabolic disorders, nutrition
- Drug discovery Cell energetics, growth and death and toxicity screening
- Cell line QC and authentication
HOW PHENOTYPE MICROARRAY TECHNOLOGY WORKS
Just as DNA Microarrays and Proteomic Technologies have made it possible to assay the level of thousands of genes or proteins all at once, Phenotype MicroArrays make it possible to quantitatively measure thousands of cellular phenotypes all at once. DNA Microarrays and Proteomic Technologies allow scientists to detect genes or proteins that are coregulated and whose patterns of change correlate with something important such as a disease state. However there is no assurance that these changes are really significant to the cell. Phenotype MicroArrays are a complementary technology providing the needed information at the cellular level.

Phenotype MicroArrays are preconfigured sets of phenotypic tests deployed on microplate panels. Each well of the array is designed to test a different phenotype after inoculation with a standardized cell suspension, allowing simultaneous testing of thousands of phenotypes in a single experiment.
Phenotype MicroArrays use Biolog’s patented redox technology, with cell respiration (NADH production) as a universal reporter. If the phenotype is strongly “positive” in a well, the cells respire actively, reducing a tetrazolium dye and forming a strong color. If it is weakly positive or negative, respiration is slowed or stopped, and less color or no color is formed.
The redox assay provides for both amplification and precise quantitation of phenotypes. Incubation and recording of phenotypic data is performed automatically by the OmniLog instrument.
To compare the phenotypes of two cell lines, one is recorded as a red tracing and one as a green tracing. These graphs can then be overlaid by the bioinformatic software to detect differences. Areas of overlap are colored yellow, whereas differences are highlighted as patches of red or green.

KINETIC DATA CAPTURE AND ANALYSIS
The OmniLog software contains a suite of algorithms that work in conjunction with the OmniLog and the Phenotype MicroArray panels to automate incubation of up to 50 microplates at a user-specified temperature with continuous collection of colorimetric assay data over time. Data analysis programs allow for display of kinetic data, management and analysis of data, and facilitates export in a variety of raw and processed forms.
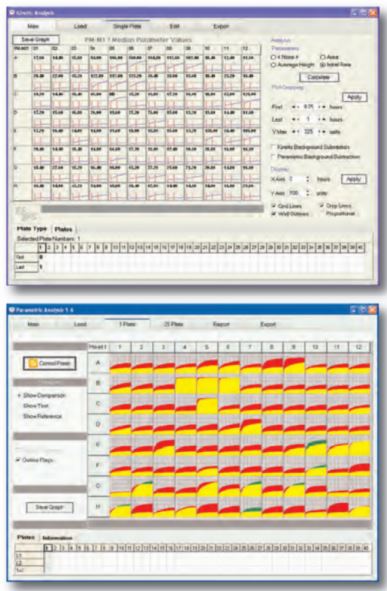
File Management/Kinetic/Parametric Analysis
- Assembles plate or PM-M panel data files into data lists
- Displays kinetic plots of the data
- Allows export of kinetic plots as bmp or jpeg files
- Extracts data lists from the File Management /Kinetic
Analysis module
- Calculates parameters from kinetic data
- Allows comparison of two data lists
- Highlights wells and generates a report on phenotypes that differ significantly in any selected kinetic analysis parameter
- Allows identification of substrates in all PM-M panels
- Links metabolic substrates in PM-M panels to the KEGG database
- Exports the original OmniLog kinetic data
- Exports data parameters for statistical and bioinformatic analysis
Click here to download the Phenotype MicroArray Mammalian Brochure
